Recent data imply that the human genome is pervasively transcribed and encodes tens of thousands of noncoding RNA (ncRNA) transcripts that play important regulatory roles in a wide array of cellular processes. Our main focus is to understand the biological functions of different classes of ncRNAs, such as microRNAs (miRNAs), long noncoding RNAs (lncRNAs), and circular RNAs (circRNAs), in human health and disease. We are currently pursuing several research projects that explore the roles of ncRNAs and RNA editing in the pathogenesis of cancer, cardiovascular disease, non-alcoholic steatohepatitis (NASH), and obesity. Our long-term goal is to translate discoveries in RNA biology into novel RNA-based therapeutic strategies for the treatment of life-threatening diseases.
THERAPEUTIC TARGETING OF METABOLIC microRNAs AS A NEW TREATMENT PARADIGM FOR NASH
NASH is a progressive subtype of nonalcoholic fatty liver disease characterized by abnormal fat accumulation in the liver, inflammation, and fibrosis. NASH is strongly associated with obesity and the metabolic syndrome and is rapidly becoming the leading cause of end-stage liver disease, liver transplantation, and hepatocellular carcinoma, underscoring the need for new therapies for treatment of NASH. This project focuses on developing miRNA-targeted therapeutics as a new treatment paradigm for NASH. Our goal is to discover drugs for effective and safe inhibition of metabolic miRNAs and assess their therapeutic potential to treat NASH in highly relevant preclinical models of NASH. Furthermore, we are evaluating hepatic and circulating miRNAs in human samples as biomarkers for NASH. We envision that the development of combined diagnostics and miRNA-targeted therapeutics has the potential to guide new treatment options for patients with NASH.
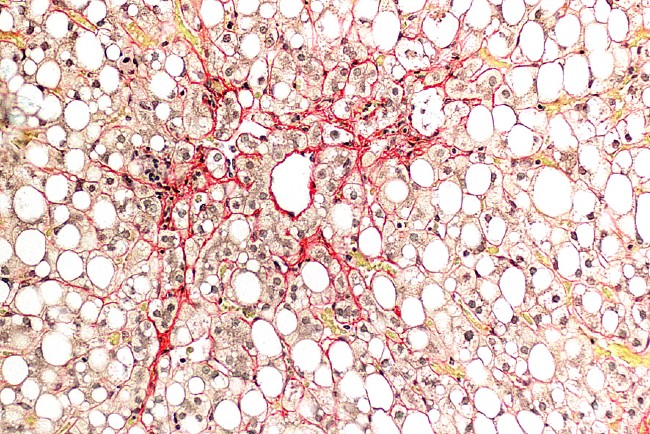
LONG NONCODING RNAs IN HEPATOCELLULAR CARCINOMA
Hepatocellular carcinoma (HCC) is the second leading cause of cancer-related deaths globally. Despite recent progress in development of HCC therapies, the median overall survival of advanced stage HCC patients is still only 12 months, which underscores the unmet medical need for novel therapeutic approaches for treatment of HCC. We aim to identify HCC-associated lncRNAs and circRNAs by RNA sequencing of HCC biopsies, uncover their molecular functions and evaluate the efficacy of design-optimized antisense oligonucleotide compounds targeting candidate lncRNAs in mouse models of HCC.
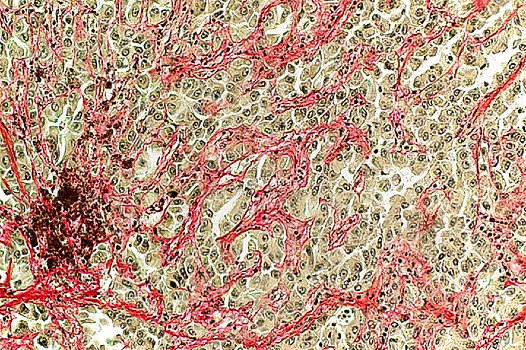
THE BIOLOGICAL ROLE OF LONG NONCODING RNAs IN CARDIOVASCULAR DISEASE
Although lncRNAs have been of great interested in the research community, their functions in cardiovascular disease are largely unknown. Using bioinformatics to generate novel hypotheses followed by in vitro and in vivo experiments in human cells and animal models, we have identified novel lncRNAs (e.g., Airn, LINC00324, LOC400043, Myolinc) and are currently elucidating their functions in the heart. Among the lncRNAs, one in particular is interesting, as we were the first team to identify an additional function of the well-known paternally imprinted lncRNA gene, Airn. Using cardiomyocytes and mouse models, we demonstrated that: (i) Airn is highly expressed in the heart; (ii) it is preferentially localized in the nucleus; (iii) silencing of Airn augments vulnerability to cell death and reduces cell migration; (iv) its expression is down-regulated following myocardial infarction; and (v) Airn binds to the RNA-binding protein Igf2bp2 and controls translation of Igf2bp2 and other genes (e.g. biglycan, Inhba) (Circulation Research, 2018).
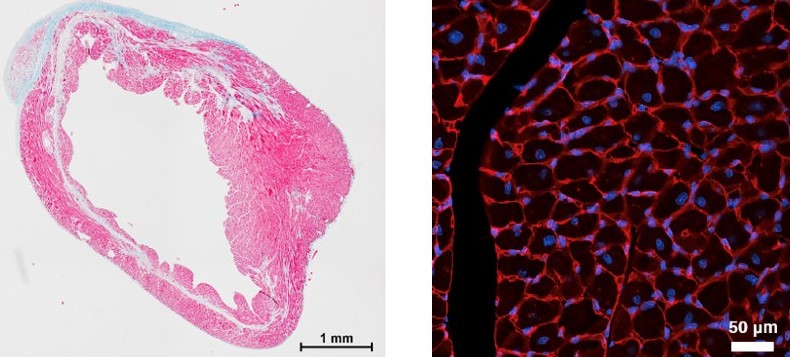
NONCODING RNAs IN MULTIPLE MYELOMA
Multiple myeloma (MM) is a hematological malignancy characterized by the accumulation of malignant plasma cells in the bone marrow leading to tumor growth, anemia, immune suppression, and myeloma bone disease. We are interested in understanding the role of ncRNAs in the pathogenesis of MM and myeloma bone disease. Previous studies have shown that miR-138 is a negative regulator of osteogenic differentiation of mesenchymal stromal cells (MSCs) and inhibiting its function enhances bone formation. We have explored the role of miR-138 in myeloma bone disease and showed that inhibition of miR-138 results in enhanced osteogenic differentiation of MM-MSCs in vitro and increases the number of endosteal osteoblastic lineage cells and bone formation rate in mouse models of myeloma bone disease. In addition, we are investigating the biological functions of selected lncRNAs associated with the pathogenesis of MM.
THE ROLE OF RNA EDITING IN HUMAN DISEASE
The recent emergence of epitranscriptomics provides an avenue for identifying RNA modifications implicated in the pathophysiology of human disease. To date, over 140 RNA modifications have been identified. These modifications are important because they can affect the fate of RNAs, including their decay, maturation, splicing, stability, and translational efficiency. Although RNA modifications have been reported in many tissues and disease contexts, detailed functional studies in the heart and cardiovascular disease (CVD) are still lacking.
A-to-I RNA editing occurs via RNA editing enzymes called "adenosine deaminases acting on RNA" (ADARs). These enzymes convert adenosine in double-stranded RNA to inosine. We have discovered that cathepsin S (CTSS), which encodes a cysteine protease associated with angiogenesis and atherosclerosis, is highly edited. Such RNA editing enables the recruitment of stabilizing RNA-binding protein HuR to the 3’-UTR of CTSS transcripts, thereby controlling CTSS mRNA stability and expression. Interestingly, ADAR1 levels and the extent of CTSS RNA editing are associated with changes in CTSS levels in patients with atherosclerotic vascular disease (Nature Medicine, 2016).
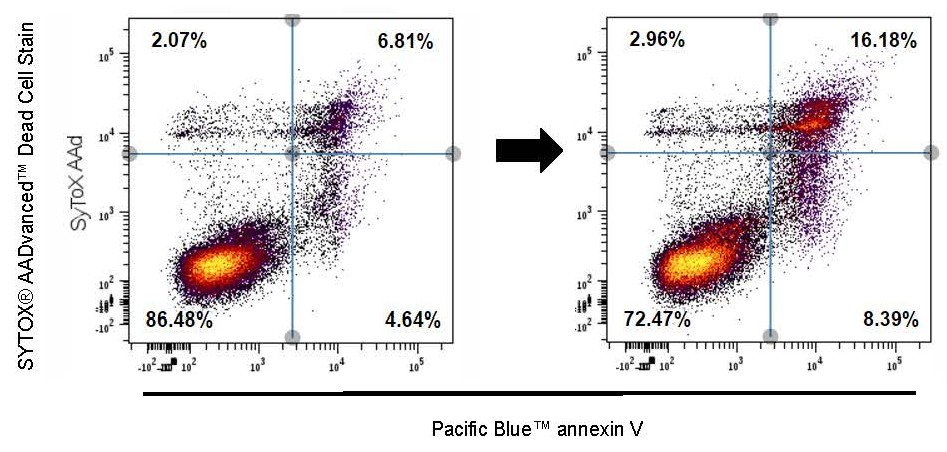
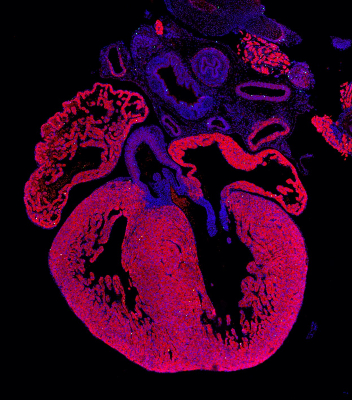
Genetic ablation of Adar1 or Adar2 results in embryonic or postnatal lethality, respectively, in mice. Adar1 null mice exhibit normal development up to E10.5, but display enhanced cellular apoptosis in multiple tissues, including the heart, and late-stage embryonic death. It remains unclear, however, whether the failure to thrive is due to reduced generation during development and/or cardiomyocyte survival. By genetically ablating Adar1 in developing cardiomyocytes, we demonstrated for the first time that Adar1 has important functions in developing hearts by controlling cardiomyocyte survival and proliferation (Circulation Research, 2020).
TOOL DEVELOPMENT
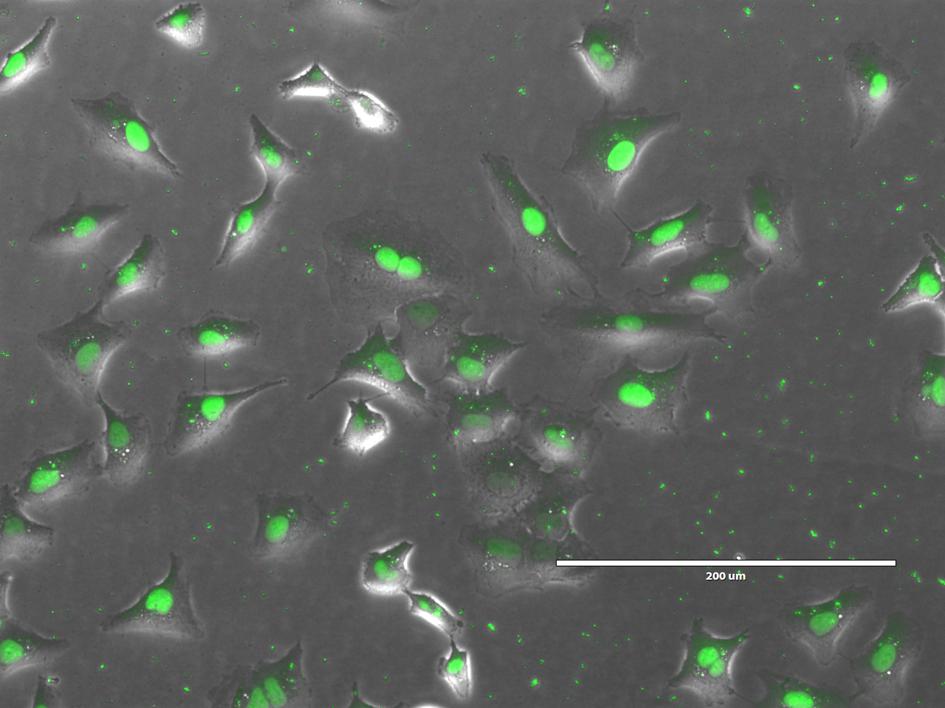
Antisense oligonucleotides (ASOs) have emerged as useful tools for knockdown of ncRNAs and hold great promise as therapeutics for the treatment of a wide range of human diseases. Most ASOs are chemically modified with phosphorothioate (PS) backbone linkages and different sugar modifications such as locked nucleic acids (LNAs) to enhance the binding affinity, stability and pharmacokinetic properties of ASOs. We have developed a tool for designing LNA-modified ASOs for ncRNA knockdown in cultured cells and in vivo. Our algorithm utilizes two ASO design paradigms with different mechanisms of action. ASOs promoting degradation of long noncoding RNA transcripts are designed as RNase H-recruiting LNA-modified PS gapmers with a central DNA gap flanked by LNA wings at the 5’ and 3’ end of the gapmer. ASOs for inhibiting miRNA function (antimiRs) are designed as LNA/DNA mixmers or short fully LNA-modified oligonucleotides with a complete PS backbone.
To address the growing need for computational approaches in RNA biology and biomedical research, we have developed a suite of bioinformatic tools. The following computational tools have been made available for researchers worldwide:
“ANGIOGENES” (http://angiogenes.uni-frankfurt.de) for expression profiles of transcripts in endothelial cells (Scientific Reports, 2016).
“C-It” (http://c-it.mpi-bn.mpg.de) for evolutionary-conserved, tissue-enriched, uncharacterized genes (Bioinformatics, 2010).
“C-It-Loci” (http://c-it-loci.uni-frankfurt.de) for evolutionary-conserved, tissue-enriched, protein-coding genes and lncRNAs (Bioinformatics, 2015).
“DRETools” (https://dretools.bitbucket.io) for studying differential RNA editings (F1000Research, 2018).
“Exon Array Analyzer (EAA)” (http://eaa.mpi-bn.mpg.de/) for analyzing Affymetrix GeneChip Exon 1.0 ST Arrays (Bioinformatics, 2009).
“FibroDB” (https://rnamedicine.shinyapps.io/FibroDB/) for expression profiles of transcripts in fibroblasts and during fibrosis (Non-Coding RNA, 2022).
“Gene Array Analyzer (GAA)” (http://gaa.mpi-bn.mpg.de/) for analyzing Affymetrix GeneChip Gene 1.0 ST Arrays (Nucleic Acids Research, 2012).
“noncoder” (http://noncoder.mpi-bn.mpg.de/) for quantifying the expression changes of lncRNAs and protein-coding genes from Affymetrix GeneChip Exon 1.0 ST Arrays (Nucleic Acids Research, 2013).
“RenalDB” (http://renaldb.uni-frankfurt.de) for in silico screening of enriched/specific transcripts of humans, mice, and zebrafish with respect to nephrotic tissues and cells, developmental stages, and other metadata (Briefings in Bioinformatics, 2018).
“RNAEditor” (http://rnaeditor.uni-frankfurt.de/) for analyzing RNA editing sites and islands from RNA-seq data (Briefings in Bioinformatics, 2017).
“SNP4Disease” (http://snp4disease.mpi-bn.mpg.de) for disease-related and/or -suspected SNPs.
“UGAHash” (http://ugahash.uni-frankfurt.de) for mapping human genomic features among various bioinformatics databases and versions of databases (Briefings in Bioinformatics, 2017).
“Vectools” (https://vectools.bitbucket.io) for working with vectors, matrices, and tables to perform statistics and machine learning analysis (F1000Research, 2018).